La epigenética estudia los cambios de expresión génica que no implican alteraciones en la secuencia de ADN. El prefijo epi- significa «encima de» o «además de», por lo tanto, la palabra epigenética engloba a todos aquellos procesos por encima de la genética y que modifican la expresión génica.
La metilación del ADN

La adición enzimática de grupos metilo (CH3-) a la molécula de ADN se denomina metilación del ADN. En mamíferos, los grupos metilo se agregan casi exclusivamente a las citosinas en los dinucleótidos CpG. Existen regiones genómicas con un elevado contenido de CpGs y se denominan islas CpG.
Se han identificado cambios anormales en la metilación del ADN en muchas enfermedades diferentes como cáncer, enfermedades inflamatorias, autoinmunes, psiquiátricas, cardiovasculares y relacionadas con la edad. Cuando estas islas CpG se hallan en las regiones promotoras del gen, la metilación del ADN actúa reprimiendo la transcripción génica.

Tratamiento con bisulfito sódico
Durante la replicación del ADN, la enzima ADN Metil-Transferasa (DNMT) re-sintetiza los grupos metilo en la cadena recién replicada. De tal forma que si realizamos una PCR, estas enzimas no estarán presenten en la reacción y por lo tanto el locus amplificado perderá la información referente al estado de metilación. Para analizar el patrón de metilación de las citosinas dentro del locus de interés, la información sobre qué citosinas están metiladas debe conservarse antes de realizar la amplificación por PCR. El tratamiento con bisulfito de sodio desamina las citosinas no metiladas en uracilos y deja intactas las citosinas metiladas (en otras palabras, las citosinas metiladas son resistentes a las modificaciones inducidas por el bisulfito de sodio).
La temperatura de fusión del ADN
La disociación de la doble hélice de ADN se denomina fusión de ADN. La temperatura de fusión del ADN depende del número de enlaces de hidrógeno que contienen las cadenas complementarias. Entre adenina (A) y timina (T) hay dos enlaces de hidrógeno mientras que entre guanina (G) y citosina (C) se forman tres enlaces. Por lo tanto, cuantas más G y C haya presentes en la secuencia, se necesitará una temperatura más alta para fundir un fragmento de ADN.
Como detectar diferencias en la temperatura de fusión
El método más utilizado para determinar la temperatura de fusión de un producto de PCR es someterlo a un gradiente de temperatura en presencia de un colorante intercalante. Estos agentes intercalantes emiten luz al unirse al ADN de doble cadena.

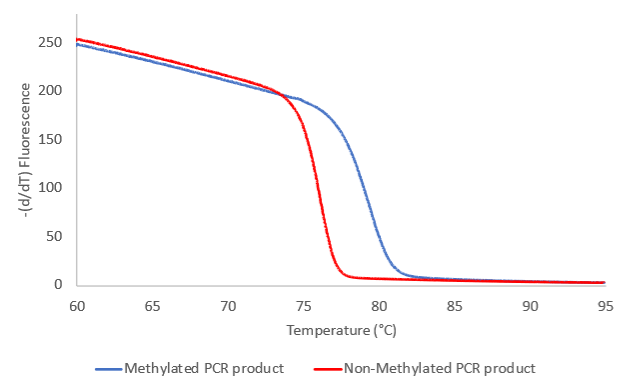
En la izquierda se muestra el resultado de los cambios de fluorescencia emitidos. Al inicio del experimento, a temperatura bajas, la fluorescencia es elevada y por lo tanto la mayoría del producto de PCR es bicatenario (A). A medida que vamos aumentando la temperatura, la fluorescencia emitida decrece lentamente, hasta una temperatura en que se rompen de forma drástica los enlaces de hidrógeno (B). A una temperatura alta no existen productos de PCR de doble cadena en la muestra y los niveles de fluorescencia están cerca de 0 (C).
La temperatura a la que observamos la fuerte caída de la fluorescencia depende del número de bandas de hidrógeno en el producto de PCR analizado y, por lo tanto, es específica del fragmento analizado.
Cómo analizar los resultados por High Resolution Melting
En principio, después de la amplificación por PCR, se obtiene un producto de PCR rico en C si el locus amplificado estaba metilado (ver gráfico a la derecha) y un producto de PCR rico en T si el locus no estaba metilado. Estos productos de PCR mostrarán diferentes perfiles de fusión. El producto de PCR amplificado de la versión no metilada del locus específico tendrá una temperatura de fusión relativamente baja y se fundirá antes en el gradiente de temperatura que el producto de PCR amplificado a partir de la versión metilada del mismo locus.
Presencia de controles en experimentos de detección de metilación
Siguiendo unas buenas prácticas de laboratorio, cualquier experimento debe incluir sus respectivos controles. En los experimentos de detección de metilación basados en PCR, se necesitan dos tipos de controles: la muestra metilada (control positivo) y la no metilada (control negativo). Esos controles son normalmente muestras de ADN químicamente modificadas que consisten en versiones metiladas y no metiladas del locus de interés y son puntos de referencia para el análisis de una muestra desconocida.

En principio, se observa hipometilación si el perfil de fusión de una muestra no se superpone con el perfil de metilación del control positivo y la hipermetilación se observa cuando el perfil de fusión de la muestra no se superpone con el perfil de fusión del control no metilado.
Methyldetect ofrece pruebas para observar si hay o no metilación para un amplio catálogo de genes. Además, si su objeto de estudio es un gen diferente de los de la lista, podemos diseñar el kit a medida.
Tanto para pedir un gen diferente como para pedir mas información o un presupuesto, puedes hacerlo en el siguiente formulario:






Un comentario en “Ensayos de metilación del ADN”
Belarus Tractor Price
Informative article very good approach towards data you provides good work. May all your dreams come true and may you find happiness and success in all your endeavors. Best wishes to you always.